Regular expressions for replication
9 minute read
As part of the publication process for my recent article on how states preempt separatist conflict, I needed to submit replication materials to the journal. I took my graduate quantitative methods sequence with the late Tom Carsey, so I’ve long been a proponent replicability efforts in social science. I also had an hourly job in grad school replicating quantitative results for multiple political science journals, so I’m very familiar with best practices for replication. Unfortunately, in the four years since I wrote the first line of code for this project, somewhere in between defending my dissertation and starting a new job (ok, fine, almost immediately after writing that first line of code), I got a little lazy.
Sometimes it’s faster (easier) to just write code that works for you, on your system, without any consideration for some poor researcher who may try to replicate your results in the future.1 This tendency was especially bad for this project because at various points in time I was writing code to run on my personal laptop and two different high performance computing clusters. This is a recipe for code that doesn’t travel well and will almost certainly fail to replicate.
There were a lot of changes I made to my code to ensure my results replicate, but the most tedious (and time consuming, by far) was cleaning up my file paths. Due to the computationally intensive GIS work and Bayesian statistics involved in the project, I ran lots of code on a cluster, and then pulled the results back to my laptop to summarize and create figures. This unsurprisingly resulted in a huge mess when looking at the project as a whole, rather than any individual script. Luckily, R and Rstudio made things (relatively) painless to fix.
File paths
Anytime you load a dataset into R, you need to specify the path to that file. The same’s true when you save R output to a file. This article started as a chapter of my dissertation, so all of the code originally lived in the Dissertation folder on my laptop. However, as I started adapting it to an article length manuscript, I created a new Conflict Preemption folder in my Projects folder. By the time the article was accepted, I had two main folders I needed to combine:
/Users/Rob/Dropbox/UNC/Dissertation/Onset/Users/Rob/Dropbox/WashU/Projects/Conflict Preemption
Both of these folders live in my Dropbox, but that’s about where the similarities end. I wrote most of the code for running models while still at UNC, so when I added new scripts to run models to respond to reviewer comments, I still stuck them in the UNC folder. That also means that all of the output of these models ended up in the UNC folder when it got transferred from the cluster. However, when I needed to do something simpler like create a time series plot of the number of separatist groups in existence, I wrote that code in the WashU folder. I also had a script in the WashU folder to load all of the results and generate plots from them. Because this script and the data it needed to load where in completely different directories, this is what I had to do to load the data to create one of the main figures:
load('/Users/Rob/Dropbox/UNC/Dissertation/Onset/Figure Data/marg_eff_pop_df_cy.RData')
Not particularly likely to work on anyone else’s computer. To fix this, I needed to move all of the data to the Conflict Preemption folder, which was easy, and then rewrite all of the code the referenced file locations, which was less easy.
Here
As a first step, I needed to chop off /Users/Rob/Dropbox/UNC/Dissertation/Onset/ from the start of every file path. All the files for the article, including both the R scripts and the various data files, now live in /Users/Rob/Dropbox/WashU/Projects/Conflict Preemption, but all of the file paths in the scripts still start with /Users/Rob/Dropbox/UNC/Dissertation/Onset, because that’s where all the files were before. You can do this just using the standard find and replace functionality built into RStudio. However, there’s no guarantee that someone in the future will correctly set R’s working directory before running the code. I used the here R package to ensure that R can always find everything it needs for my code. All you have to do is wrap file paths in the here() function in the package, and they’ll be automatically completed with the full file path, letting R find your files.2
You need to use the relative path to each file, so for a file with an absolute path of /Users/Rob/Dropbox/WashU/Projects/Conflict Preemption/Figure Data/marg_eff_pop_df_cy.RData, the relative path (relative to the project folder of /Users/Rob/Dropbox/WashU/Projects/Conflict Preemption) would be Figure Data/marg_eff_pop_df_cy.RData. The final bit of R code looks like this:
load(here('Figure Data/marg_eff_pop_df_cy.RData'))
The addition of that here() in between load() and the file path means that things are no longer as simple as finding and replacing the start of the file path.
Regular expressions
Luckily, I was able to take advantage of RStudio’s built in support for regular expressions to save myself from having to manually change each line of code that either loaded or saved a file. Regular expressions are a powerful way to search through and manipulate text. You can activate them in RStudio’s find and replace dialog by checking the Regex box:
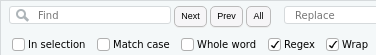
Once you’ve done that, certain characters in your search will no longer be interpreted literally. The most important difference is probably ., which is a stand-in for any character.3 This is similar to how * is a wildcard in the Unix shell, e.g., you can use ls *.R to list all R script files in a folder. The main regular expression feature I used is the capturing group, which allows you to identify and extract a subset of a line of text. You designate a capturing group by surrounding the desired text with parentheses. To fix all of the code loading RData files from the Figure Data folder, my regular expression looked like this:
'/Users/Rob/Dropbox/UNC/Dissertation/Onset/(Figure Data/.*\.RData)'
It starts with /Users/Rob/Dropbox/UNC/Dissertation/Onset/, which is the part I want to get rid of. Next, (Figure Data/.*\.RData)' tells the regular expression to look for any character (.) repeated an unlimited number of times (*) followed by .RData. Because . is a special character in regular expressions, we have to escape it with a backslash (\). This will match any file name ending in .RData in the Figure Data folder. If we left out the leading /Users/Rob/Dropbox/UNC/Dissertation/Onset/, we end up with the capturing group we want, but since /Users/Rob/Dropbox/UNC/Dissertation/Onset/ wouldn’t be part of the search string, it wouldn’t end up getting replaced. This is the same reason we need to include the opening and closing quotation marks; if we didn’t, we’d end up with a here() command inside quotation marks, which R would just treat as a string and not a command.
At this point I had the core of the line that I wanted to keep, but now I needed to extract it and place it inside of a call to here(). You accomplish this goal using a backreference to the capturing group. To reference the first capturing group, you use either \1 or $1 depending on which version of regular expressions you are using. This is often very difficult to figure out, and is one of the most annoying things about regular expressions. You’ll often just have to experiment and find out which one to use through trial and error. Luckily RStudio accepts either version!
To replace the absolute path with a relative one wrapped in a here() call, this is what I typed into the Replace field in the find and replace dialog:
here('$1')
and it resulted in this:
here('Figure Data/marg_eff_pop_df_cy.RData')
Thanks to the power of capture groups, you can just hit the replace all button and instantly transform every file path into a much more portable and replication-friendly one.
A little bit faster now
If you’re feeling really confident that you moved every file correctly, you can replace all file paths with the following regular expression:
'/Users/Rob/Dropbox/UNC/Dissertation/Onset/(.*\..*)'
This will get any files with file extensions (the \. followed by .* to ensure there’s at least one character after a literal period), as well as any preceding subdirectories (the initial .*) and stick them all into the resulting here() call. As an example, this will successfully turn this: fileConn <- file(here::here(‘Tables/pd_pop_cy.tex’))
groups <- readRDS('/Users/Rob/Dropbox/Dissertation/Onset/Input Data/groups_nightlights.RDS')
into this:
groups <- readRDS(here::here('Input Data/groups_nightlights.RDS'))
I’m using ‘replication’ here to mean that the code used to generate quantitative results from a dataset should produce those same results when run by another researcher, not in the sense that means that independent researchers following the published protocol can collect the data themselves and arrive at the same conclusion. I use the term ‘reproducible’ to describe this property. Annoyingly, different fields use opposing definitions of these two terms. ↩
Specifically,
here()will key into the.Rprojfile included in my replication materials and use that to properly locate everything else. ↩Except for newlines, carriage returns, and other end of line special characters. ↩
